Running species#
This tutorial shows what a typical workflow with species looks like. We will compare the colors and magnitudes of a few directly imaged companions with those of field dwarfs from the Database of Ultracool Parallaxes.
Initiating species#
We start by importing the species toolkit.
[1]:
from species import SpeciesInit
from species.data.database import Database
from species.read.read_color import ReadColorMagnitude
from species.plot.plot_color import plot_color_magnitude
Next, the configuration file (species_config.ini) is read from the working folder. Since the file does not exist, it is created with default values for the path of the HDF5 database and the location where the external data is stored. The database and data folder are also created since this is the first time that we run species with the current configuration file.
[2]:
SpeciesInit()
==============
species v0.8.0
==============
Working folder: /Users/tomasstolker/applications/species/docs/tutorials
Creating species_config.ini... [DONE]
Creating species_database.hdf5... [DONE]
Creating data folder... [DONE]
Configuration settings:
- Database: /Users/tomasstolker/applications/species/docs/tutorials/species_database.hdf5
- Data folder: /Users/tomasstolker/applications/species/docs/tutorials/data
- Magnitude of Vega: 0.03
Multiprocessing: mpi4py installed
Process number 1 out of 1...
[2]:
<species.core.species_init.SpeciesInit at 0x15695d510>
Adding a photometric library#
A Database object is now created, which can be used for importing data from the chosen data folder into the database.
[3]:
database = Database()
The photometric data and parallaxes from the Database of Ultracool Parallaxes are downloaded and added to the database with the add_photometry method of Database.
[4]:
database.add_photometry('vlm-plx')
Downloading data from 'https://home.strw.leidenuniv.nl/~stolker/species/vlm-plx-all.fits' to file '/Users/tomasstolker/applications/species/docs/tutorials/data/vlm-plx-all.fits'.
-----------------------
Add photometric library
-----------------------
Database tag: vlm-plx
Library: Database of Ultracool Parallaxes
100%|████████████████████████████████████████| 314k/314k [00:00<00:00, 448MB/s]
Adding companion data#
A library of magnitudes and parallaxes of directly imaged companions are available in the companion_data file. These can be added to the Database with the add_companion method (alternatively, photometric data of individual objects can be manually imported with add_object). In this example, we add the distance and magnitudes of 51 Eri b, beta Pic b, and PZ Tel B to the database. This will automatically download the filter profiles and a flux calibrated spectrum of Vega. These are required for the flux calibration of the companion magnitudes.
[5]:
database.add_companion('51 Eri b', verbose=False)
database.add_companion('beta Pic b', verbose=False)
database.add_companion('PZ Tel B', verbose=False)
Add companion: ['51 Eri b']
Downloading data from 'https://archive.stsci.edu/hlsps/reference-atlases/cdbs/current_calspec/alpha_lyr_stis_011.fits' to file '/Users/tomasstolker/applications/species/docs/tutorials/data/alpha_lyr_stis_011.fits'.
100%|████████████████████████████████████████| 288k/288k [00:00<00:00, 121MB/s]
Adding spectrum: Vega
Reference: Bohlin et al. 2014, PASP, 126
URL: https://ui.adsabs.harvard.edu/abs/2014PASP..126..711B/abstract
Add companion: ['beta Pic b']
/Users/tomasstolker/applications/species/species/data/database.py:1373: UserWarning: Found 33 fluxes with NaN in the data of GPI_YJHK. Removing the spectral fluxes that contain a NaN.
warnings.warn(
Add companion: ['PZ Tel B']
Reading color-magnitude data#
The colors and magnitudes from the Database of Ultracool Parallaxes are read from the database in the working folder by first creating an instance of ReadColorMagnitude.
[6]:
colormag = ReadColorMagnitude(library='vlm-plx',
filters_color=('MKO/NSFCam.J', 'MKO/NSFCam.H'),
filter_mag='MKO/NSFCam.J')
--------------------
Read color-magnitude
--------------------
Database tag: vlm-plx
Library type: phot_lib
Filters color: ('MKO/NSFCam.J', 'MKO/NSFCam.H')
Filter magnitude: MKO/NSFCam.J
The data of the field dwarfs are now read with the get_color_magnitude method. This returns a ColorMagBox with the requested data.
[7]:
colorbox = colormag.get_color_magnitude(object_type='field')
-------------------
Get color-magnitude
-------------------
Object type: field
Returning ColorMagBox with 241 objects
We also create a list with the color and magnitude filter names that we want to extract for the directly imaged companions.
[8]:
objects = [('51 Eri b', 'MKO/NSFCam.J', 'MKO/NSFCam.H', 'MKO/NSFCam.J'),
('beta Pic b', 'Paranal/NACO.J', 'Paranal/NACO.H', 'Paranal/NACO.J'),
('PZ Tel B', 'Paranal/NACO.J', 'Paranal/NACO.H', 'Paranal/NACO.J')]
Plotting a color-magnitude diagram#
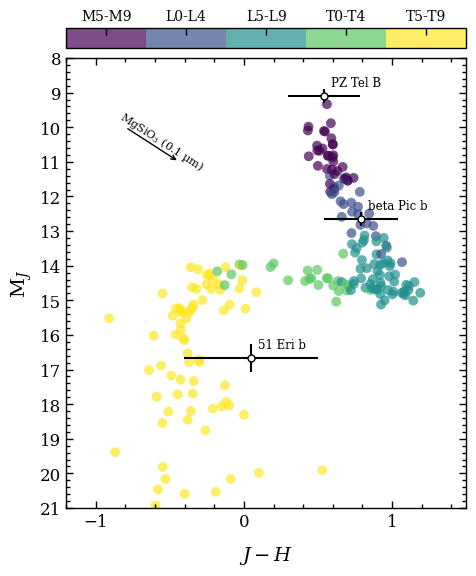
We will now combine the selected data in a color-magnitude diagram. The ColorMagBox is provided in a list to the plot_color_magnitude function. The list with directly imaged objects is provided as argument of objects. We also add a reddening vector to show how the color changes for a size
distribution of enstatite grains with a geometric mean radius of 0.1 \(\mu\)m and an extinction in the \(J\) band of 1 mag. The discrete colorbar shows the spectral types of the field dwarfs and the directly imaged objects are labeled individually.
[9]:
fig = plot_color_magnitude(boxes=[colorbox, ],
objects=objects,
reddening=[(('MKO/NSFCam.J', 'MKO/NSFCam.H'), ('MKO/NSFCam.J', 1.), 'MgSiO3', 0.1, (-0.8, 10.))],
label_x=r'$J - H$',
label_y=r'M$_J$',
xlim=(-1.2, 1.5),
ylim=(21., 8.),
offset=(-0.08, -0.08),
companion_labels=True,
field_range=('late M', 'late T'),
legend=None,
output=None)
----------------------------
Plot color-magnitude diagram
----------------------------
Boxes:
- ColorMagBox
Objects:
- 51 Eri b: ('MKO/NSFCam.J', 'MKO/NSFCam.H', 'MKO/NSFCam.J')
- beta Pic b: ('Paranal/NACO.J', 'Paranal/NACO.H', 'Paranal/NACO.J')
- PZ Tel B: ('Paranal/NACO.J', 'Paranal/NACO.H', 'Paranal/NACO.J')
Spectral range: ('late M', 'late T')
Companion labels: True
Accretion markers: False
Mass labels: None
Teff labels: None
Reddening:
- (('MKO/NSFCam.J', 'MKO/NSFCam.H'), ('MKO/NSFCam.J', 1.0), 'MgSiO3', 0.1, (-0.8, 10.0))
Figure size: (4.0, 4.8)
Limits x axis: (-1.2, 1.5)
Limits y axis: (21.0, 8.0)
Downloading optical constants (87 kB)... [DONE]
Unpacking optical constants... [DONE]
Adding optical constants of MgSiO3... [DONE]
Adding optical constants of Fe... [DONE]
Downloading log-normal dust cross sections (231 kB)... [DONE]
Adding log-normal dust cross sections:
- Data shape (n_wavelength, n_radius, n_sigma): (67, 20, 20)
- Wavelength range: 0.4 - 10.0 um
- Mean geometric radius range: 0.001 - 10.0 um
- Geometric standard deviation range: 1.0 - 10.0
Downloading power-law dust cross sections (231 kB)... [DONE]
Adding power-law dust cross sections:
- Data shape (n_wavelength, n_radius, n_exponent): (132, 50, 50)
- Wavelength range: 0.4 - 10.0 um
- Maximum grain radius range: 0.01 - 100.0 um
- Power-law exponent range: -10.0 - 10.0
The plot_color_magnitude function returned the Figure object of the plot. The functionalities of Matplotlib can be used for further customization of the plot. For example, the axes of the plot are stored at the axes attribute of Figure.
[10]:
fig.axes
[10]:
[<Axes: xlabel='$J - H$', ylabel='M$_J$'>, <Axes: >]
